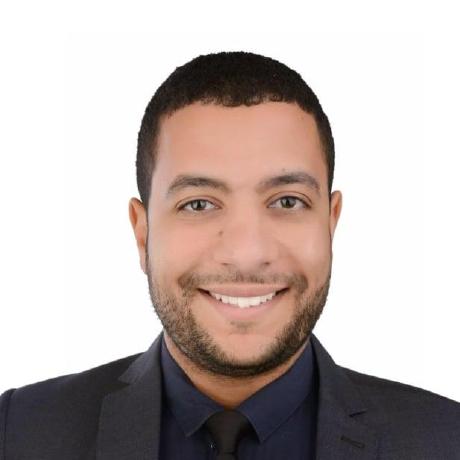
GeneticAlgorithmPython
Source code of PyGAD, a Python 3 library for building the genetic algorithm and training machine learning algorithms (Keras & PyTorch).
BSD-3-CLAUSE License
Bot releases are hidden (Show)
Published by ahmedfgad over 3 years ago
-
Issue #40 is solved. Now, the
Nonevalue works with thecrossover_typeandmutation_typeparameters: https://github.com/ahmedfgad/GeneticAlgorithmPython/issues/40 - The
gene_typeparameter supports accepting alist/tuple/numpy.ndarrayof numeric data types for the genes. This helps to control the data type of each individual gene. Previously, thegene_typecan be assigned only to a single data type that is applied for all genes. - A new
boolattribute namedgene_type_singleis added to thepygad.GAclass. It isTruewhen there is a single data type assigned to thegene_typeparameter. When thegene_typeparameter is assigned alist/tuple/numpy.ndarray, thengene_type_singleis set toFalse. - The
mutation_by_replacementflag now has no effect ifgene_spaceexists except for the genes withNonevalues. For example, forgene_space=[None, [5, 6]]themutation_by_replacementflag affects only the first gene which hasNonefor its value space. - When an element has a value of
Nonein thegene_spaceparameter (e.g.gene_space=[None, [5, 6]]), then its value will be randomly generated for each solution rather than being generate once for all solutions. Previously, the gene withNonevalue ingene_spaceis the same across all solutions - Some changes in the documentation according to issue #32: https://github.com/ahmedfgad/GeneticAlgorithmPython/issues/32
Published by ahmedfgad over 3 years ago
PyGAD 2.13.0
Release Date: 12 March 2021
- A new
boolparameter calledallow_duplicate_genesis supported. IfTrue, which is the default, then a solution/chromosome may have duplicate gene values. IfFalse, then each gene will have a unique value in its solution. Check the Prevent Duplicates in Gene Values section for more details. - The
last_generation_fitnessis updated at the end of each generation not at the beginning. This keeps the fitness values of the most up-to-date population assigned to thelast_generation_fitnessparameter.
Published by ahmedfgad over 3 years ago
Release Date: 20 February 2021
- 4 new instance attributes are added to hold temporary results after each generation:
last_generation_fitnessholds the fitness values of the solutions in the last generation,last_generation_parentsholds the parents selected from the last generation,last_generation_offspring_crossoverholds the offspring generated after applying the crossover in the last generation, andlast_generation_offspring_mutationholds the offspring generated after applying the mutation in the last generation. You can access these attributes inside theon_generation()method for example. - A bug fixed when the
initial_populationparameter is used. The bug occurred due to a mismatch between the data type of the array assigned toinitial_populationand the gene type in thegene_typeattribute. Assuming that the array assigned to theinitial_populationparameter is((1, 1), (3, 3), (5, 5), (7, 7))which has typeint. Whengene_typeis set tofloat, then the genes will not be float but casted tointbecause the defined array hasinttype. The bug is fixed by forcing the array assigned toinitial_populationto have the data type in thegene_typeattribute. Check the issue at GitHub: https://github.com/ahmedfgad/GeneticAlgorithmPython/issues/27
Thanks to Marios Giouvanakis, a PhD candidate in Electrical & Computer Engineer, Aristotle University of Thessaloniki (Αριστοτέλειο Πανεπιστήμιο Θεσσαλονίκης), Greece, for emailing me about these issues.
Published by ahmedfgad over 3 years ago
PyGAD 2.11.0
Release Date: 16 February 2021
- In the
gene_spaceargument, the user can use a dictionary to specify the lower and upper limits of the gene. This dictionary must have only 2 items with keyslowandhighto specify the low and high limits of the gene, respectively. This way, PyGAD takes care of not exceeding the value limits of the gene. For a problem with only 2 genes, then usinggene_space=[{'low': 1, 'high': 5}, {'low': 0.2, 'high': 0.81}]means the accepted values in the first gene start from 1 (inclusive) to 5 (exclusive) while the second one has values between 0.2 (inclusive) and 0.85 (exclusive). For more information, please check the Limit the Gene Value Range section of the documentation. - The
plot_result()method returns the figure so that the user can save it. - Bug fixes in copying elements from the gene space.
- For a gene with a set of discrete values (more than 1 value) in the
gene_spaceparameter like[0, 1], it was possible that the gene value may not change after mutation. That is if the current value is 0, then the randomly selected value could also be 0. Now, it is verified that the new value is changed. So, if the current value is 0, then the new value after mutation will not be 0 but 1.
Published by ahmedfgad almost 4 years ago
A bug fix when save_best_solutions=True. Refer to this issue for more information: https://github.com/ahmedfgad/GeneticAlgorithmPython/issues/25
Published by ahmedfgad almost 4 years ago
Changes in PyGAD 2.10.1
- In the
gene_spaceparameter, anyNonevalue (regardless of its index or axis), is replaced by a randomly generated number based on the 3 parametersinit_range_low,init_range_high, andgene_type. So, theNonevalue in[..., None, ...]or[..., [..., None, ...], ...]are replaced with random values. This gives more freedom in building the space of values for the genes. - All the numbers passed to the
gene_spaceparameter are casted to the type specified in thegene_typeparameter. - The
numpy.uintdata type is supported for the parameters that accept integer values. - In the
pygad.kerasgamodule, themodel_weights_as_vector()function uses thetrainableattribute of the model's layers to only return the trainable weights in the network. So, only the trainable layers with theirtrainableattribute set toTrue(trainable=True), which is the default value, have their weights evolved. All non-trainable layers with thetrainableattribute set toFalse(trainable=False) will not be evolved. Thanks to Prof. Tamer A. Farrag for pointing about that at GitHub.
Published by ahmedfgad almost 4 years ago
- Support of a new module
pygad.torchgato train PyTorch models using PyGAD. Check its documentation. - Support of adaptive mutation where the mutation rate is determined by the fitness value of each solution. Read the Adaptive Mutation section for more details. Also, read this paper: Libelli, S. Marsili, and P. Alba. "Adaptive mutation in genetic algorithms." Soft computing 4.2 (2000): 76-80.
- Before the
run()method completes or exits, the fitness value of the best solution in the current population is appended to thebest_solution_fitnesslist attribute. Note that the fitness value of the best solution in the initial population is already saved at the beginning of the list. So, the fitness value of the best solution is saved before the genetic algorithm starts and after it ends. - When the parameter
parent_selection_typeis set tosss(steady-state selection), then a warning message is printed if the value of thekeep_parentsparameter is set to 0. - More validations to the user input parameters.
- The default value of the
mutation_percent_genesis set to the string"default"rather than the integer 10. This change helps to know whether the user explicitly passed a value to themutation_percent_genesparameter or it is left to its default one. The"default"value is later translated into the integer 10. - The
mutation_percent_genesparameter is no longer accepting the value 0. It must be>0and<=100. - The built-in
warningsmodule is used to show warning messages rather than just using theprint()function. - A new
boolparameter calledsuppress_warningsis added to the constructor of thepygad.GAclass. It allows the user to control whether the warning messages are printed or not. It defaults toFalsewhich means the messages are printed. - A helper method called
adaptive_mutation_population_fitness()is created to calculate the average fitness value used in adaptive mutation to filter the solutions. - The
best_solution()method accepts a new optional parameter calledpop_fitness. It accepts a list of the fitness values of the solutions in the population. IfNone, then thecal_pop_fitness()method is called to calculate the fitness values of the population.
Published by ahmedfgad almost 4 years ago
Changes in PyGAD 2.9.0 (06 December 2020):
- The fitness values of the initial population are considered in the
best_solutions_fitnessattribute. - An optional parameter named
save_best_solutionsis added. It defaults toFalse. When it isTrue, then the best solution after each generation is saved into an attribute namedbest_solutions. IfFalse, then no solutions are saved and thebest_solutionsattribute will be empty. - Scattered crossover is supported. To use it, assign the
crossover_typeparameter the value"scattered". - NumPy arrays are now supported by the
gene_spaceparameter. - The following parameters (
gene_type,crossover_probability,mutation_probability,delay_after_gen) can be assigned to a numeric value of any of these data types:int,float,numpy.int,numpy.int8,numpy.int16,numpy.int32,numpy.int64,numpy.float,numpy.float16,numpy.float32, ornumpy.float64.
Published by ahmedfgad about 4 years ago
- Bug fix in applying the crossover operation when the
crossover_probabilityparameter is used. Thanks to Eng. Hamada Kassem, Research and Teaching Assistant, Construction Engineering and Management, Faculty of Engineering, Alexandria University, Egypt.
Published by ahmedfgad about 4 years ago
Support of a new module named pygad.kerasga to train Keras models using the genetic algorithm.
Published by ahmedfgad about 4 years ago
Bug fix to support building and training regression neural networks with multiple outputs.
Published by ahmedfgad about 4 years ago
A bug fix when the problem_type argument is set to regression.
Published by ahmedfgad about 4 years ago
Changes in PyGAD 2.7.0 (11 September 2020):
- The
learning_rateparameter in thepygad.nn.train()function defaults to 0.01. - Added support of building neural networks for regression using the new parameter named
problem_type. It is added as a parameter to bothpygad.nn.train()andpygad.nn.predict()functions. The value of this parameter can be either classification or regression to define the problem type. It defaults to classification. - The activation function for a layer can be set to the string
"None"to refer that there is no activation function at this layer. As a result, the supported values for the activation function are"sigmoid","relu","softmax", and"None".
To build a regression network using the pygad.nn module, just do the following:
- Set the
problem_typeparameter in thepygad.nn.train()andpygad.nn.predict()functions to the string"regression". - Set the activation function for the output layer to the string
"None". This sets no limits on the range of the outputs as it will be from-infinityto+infinity. If you are sure that all outputs will be nonnegative values, then use the ReLU function.
Check the documentation of the pygad.nn module for an example that builds a neural network for regression. The regression example is also available at this GitHub project: https://github.com/ahmedfgad/NumPyANN
To build and train a regression network using the pygad.gann module, do the following:
- Set the
problem_typeparameter in thepygad.nn.train()andpygad.nn.predict()functions to the string"regression". - Set the
output_activationparameter in the constructor of thepygad.gann.GANNclass to"None".
Check the documentation of the pygad.gann module for an example that builds and trains a neural network for regression. The regression example is also available at this GitHub project: https://github.com/ahmedfgad/NeuralGenetic
To build a classification network, either ignore the problem_type parameter or set it to "classification" (default value). In this case, the activation function of the last layer can be set to any type (e.g. softmax).
Published by ahmedfgad about 4 years ago
Release Date: 6 August 2020
- A bug fix in assigning the value to the
initial_populationparameter. - A new parameter named
gene_typeis added to control the gene type. It can be eitherintorfloat. It has an effect only when the parametergene_spaceisNone. - 7 new parameters that accept callback functions:
on_start,on_fitness,on_parents,on_crossover,on_mutation,on_generation, andon_stop.
Published by ahmedfgad about 4 years ago
Changes in PyGAD 2.5.0 - Release date: 19 July 2020
- 2 new optional parameters added to the constructor of the
pygad.GAclass which arecrossover_probabilityandmutation_probability.
While applying the crossover operation, each parent has a random value generated between 0.0 and 1.0. If this random value is less than or equal to the value assigned to thecrossover_probabilityparameter, then the parent is selected for the crossover operation.
For the mutation operation, a random value between 0.0 and 1.0 is generated for each gene in the solution. If this value is less than or equal to the value assigned to themutation_probability, then this gene is selected for mutation. - A new optional parameter named
linewidthis added to theplot_result()method to specify the width of the curve in the plot. It defaults to 3.0. - Previously, the indices of the genes selected for mutation was randomly generated once for all solutions within the generation. Currently, the genes' indices are randomly generated for each solution in the population. If the population has 4 solutions, the indices are randomly generated 4 times inside the single generation, 1 time for each solution.
- Previously, the position of the point(s) for the single-point and two-points crossover was(were) randomly selected once for all solutions within the generation. Currently, the position(s) is(are) randomly selected for each solution in the population. If the population has 4 solutions, the position(s) is(are) randomly generated 4 times inside the single generation, 1 time for each solution.
- A new optional parameter named
gene_spaceas added to thepygad.GAclass constructor. It is used to specify the possible values for each gene in case the user wants to restrict the gene values. It is useful if the gene space is restricted to a certain range or to discrete values.
Assuming that all genes have the same global space which include the values 0.3, 5.2, -4, and 8, then those values can be assigned to the gene_space parameter as a list, tuple, or range. Here is a list assigned to this parameter. By doing that, then the gene values are restricted to those assigned to the gene_space parameter.
gene_space = [0.3, 5.2, -4, 8]
If some genes have different spaces, then gene_space should accept a nested list or tuple. In this case, its elements could be:
- List, tuple, or range: It holds the individual gene space.
- Number (int/float): A single value to be assigned to the gene. This means this gene will have the same value across all generations.
-
None: A gene with its space set toNoneis initialized randomly from the range specified by the 2 parametersinit_range_lowandinit_range_high. For mutation, its value is mutated based on a random value from the range specified by the 2 parametersrandom_mutation_min_valandrandom_mutation_max_val. If all elements in thegene_spaceparameter areNone, the parameter will not have any effect.
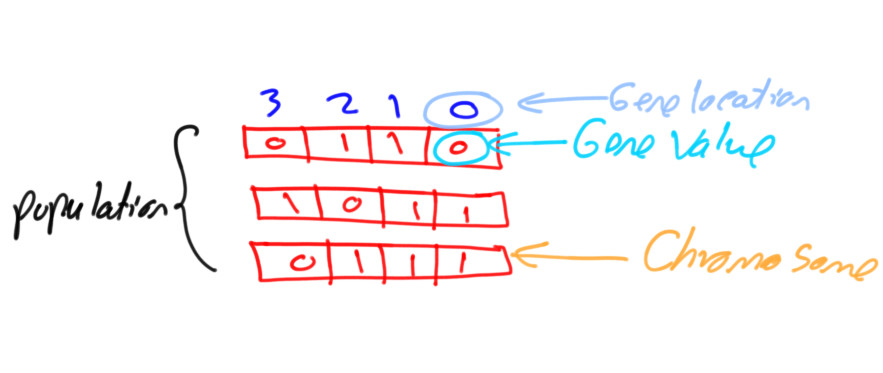
Assuming that a chromosome has 2 genes and each gene has a different value space. Then the gene_space could be assigned a nested list/tuple where each element determines the space of a gene. According to the next code, the space of the first gene is [0.4, -5] which has 2 values and the space for the second gene is [0.5, -3.2, 8.8, -9] which has 4 values.
gene_space = [[0.4, -5], [0.5, -3.2, 8.2, -9]]
For a 2 gene chromosome, if the first gene space is restricted to the discrete values from 0 to 4 and the second gene is restricted to the values from 10 to 19, then it could be specified according to the next code.
gene_space = [range(5), range(10, 20)]
If the user did not assign the initial population to the initial_population parameter, the initial population is created randomly based on the gene_space parameter. Moreover, the mutation is applied based on this parameter.
Published by ahmedfgad over 4 years ago
Changes in PyGAD 2.4.0:
- A new parameter named
delay_after_genis added which accepts a non-negative number specifying the time in seconds to wait after a generation completes and before going to the next generation. It defaults to0.0which means no delay after the generation. - The passed function to the
callback_generationparameter of the pygad.GA class constructor can terminate the execution of the genetic algorithm if it returns the stringstop. This causes therun()method to stop.
One important use case for that feature is to stop the genetic algorithm when a condition is met before passing though all the generations. The user may assigned a value of 100 to the num_generations parameter of the pygad.GA class constructor. Assuming that at generation 50, for example, a condition is met and the user wants to stop the execution before waiting the remaining 50 generations. To do that, just make the function passed to the callback_generation parameter to return the string stop.
Here is an example of a function to be passed to the callback_generation parameter which stops the execution if the fitness value 70 is reached. The value 70 might be the best possible fitness value. After being reached, then there is no need to pass through more generations because no further improvement is possible.
def func_generation(ga_instance):
if ga_instance.best_solution()[1] >= 70:
return "stop"
Published by ahmedfgad over 4 years ago
Changes in PyGAD 1.0.19 (4 May 2020):
- The attributes are moved from the class scope to the instance scope.
- Raising a
ValueErrorexception on passing incorrect values to the parameters. - Two new parameters are added (
init_rand_highandinit_rand_high) allowing the user to customize the range from which the genes values in the initial population are selected. - The code object
__code__of the passed fitness function is checked to ensure it has the right number of parameters.